class: inverse, bottom background-image: url(https://plus.unsplash.com/premium_photo-1686050878751-89499d28d153?ixlib=rb-4.1.0&ixid=M3wxMjA3fDB8MHxzZWFyY2h8MXx8bmlnaHRza3l8ZW58MHx8MHx8fDA%3D&auto=format&fit=crop&q=60&w=500) background-size: cover # .Large[{ggdims}] ## .small[Toward a viz- and code-first hitchhiker’s guide to dimensionality reduction] ## .small[github.com/EvaMaeRey/ggdims] #### .tiny[Gina (Evangeline) Reynolds | 2025-11-14 | ASA, COWY, Image credit: Ales Krivec, Upsplash] <style type="text/css"> /* Change default body text size */ .remark-slide-content { font-size: 32px; /* Or use em, rem, or viewport units like 2vw */ } /* Change font size for a specific class (e.g., for larger text) */ .large-text { font-size: 1.5em; } /* Change font size for code blocks */ .remark-code { font-size: 0.9em; } /* Change font size for inline code */ .remark-inline-code { font-size: 0.9em; } </style> --- ### cliffnotes for talk --- This pattern for ggplot2 extension ... -- ``` r library(ggplot2) ggplot(data = my_high_dimensional_data) + aes(dims = dims(var1:var200, var205)) + geom_dim_red_technique() # geom_pca(), geom_tsne ``` -- ... isn't a pattern* ... -- And maybe it should be. -- \* or is it and I don't know that? --- `aes(dims = dims(var1:var200, var205))`  --- class: inverse, middle, center {ggdims} explores viz- and code- first dimensionality reduction -- - ~~A 3-week-old github repo/package~~ -- Rest of talk: how we got here (a longer time coming) --- class: inverse, middle, center # ggplot2 (2007) lets you intuitively translate variables to visual representation. --- `gplot(my_data) +` `aes(x = age) + ` `aes(color = sex) +` `aes(size = employment status) +` `geom_line()` <!-- --- --> <!-- class: inverse, middle --> <!-- # You specify how variables (e.g. sex, age, employment status) ... --> <!-- -- --> <!-- # ... are to be communicated via visual channels (x position, y position, color, transparency, etc). --> --- class: inverse # But... ggplot2 allows individual-variable- -- to- individual-visual-channel mapping --- class: inverse # A problem with 1-to-1: -- ## Doesn't lend itself easily to visualizations in the world of dimension reduction (e.g. PCA, t-SNE, UMAP) ??? ## However, in ggplot2 these specifications are individual-variable-to-individual-visual-channel which does not lend itself easily to visualizations in the world of dimension reduction (e.g. PCA, T-sne, UMAP). --- ``` r library(ggplot2) ggplot(data = my_high_dimensional_data) + aes(dims = dims(var1:var200, var205)) + # << geom_dimred_technique() # e.g. geom_pca(), geom_tsne(), geom_umap() ``` --- > "You really should 'algorithm' first, and then use the results in a data-viz." -- (algorithm used as a verb here) --- # 2016 https://distill.pub/2016/misread-tsne/ 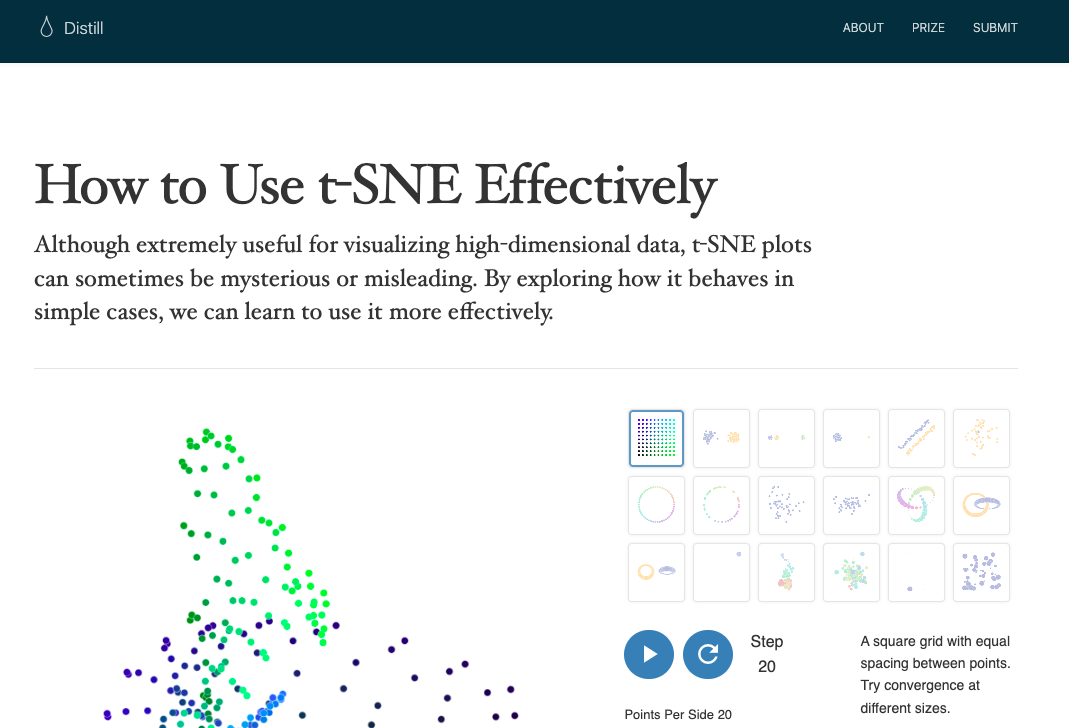 ---  --- Martin Wattenberg 'How to use t-SNE effectively' 2016: > Paraphrase: 'it's worth taking the time to get a feel for what the algorithm does with some familiar, well understood distributions' -- > Further paraphrase: Jump right in. Do some dimensionality reduction. Visualize. ---  --- > "But maybe Wattenberg got it wrong..." --- Leland McInnes "Bluffer's Guide" > "I'm gonna try and give as much of a whirlwind tour of the entire field of dimention reduction as I can in the time available."  ---  -- > 'The similarities are greater than the differences' ?... --- > Shouldn't we be able to visually fly (hitchhike) through trying out some of these techniques? -- > Should we have to learn a different paradigm/syntax for every technique? -- > What if the syntax were something already familiar (2D viz - ggplot2 scatterplot)? --- count: false .panel1-scatter-auto[ ``` r *library(ggplot2) ``` ] .panel2-scatter-auto[ ] --- count: false .panel1-scatter-auto[ ``` r library(ggplot2) # data *ggplot(mtcars) ``` ] .panel2-scatter-auto[ <!-- --> ] --- count: false .panel1-scatter-auto[ ``` r library(ggplot2) # data ggplot(mtcars) + # variable mapping * aes(x = disp, * y = mpg) ``` ] .panel2-scatter-auto[ <!-- --> ] --- count: false .panel1-scatter-auto[ ``` r library(ggplot2) # data ggplot(mtcars) + # variable mapping aes(x = disp, y = mpg) + # layer * geom_point() ``` ] .panel2-scatter-auto[ <!-- --> ] <style> .panel1-scatter-auto { color: black; width: 38.6060606060606%; hight: 32%; float: left; padding-left: 1%; font-size: 80% } .panel2-scatter-auto { color: black; width: 59.3939393939394%; hight: 32%; float: left; padding-left: 1%; font-size: 80% } .panel3-scatter-auto { color: black; width: NA%; hight: 33%; float: left; padding-left: 1%; font-size: 80% } </style> --- ``` r ggplot(mtcars) + # data aes(????) # var mapping geom_pca() # layer - options inside parentheses ggplot(mtcars) + # data aes(????) # var mapping geom_tsne() # layer ggplot(mtcars) + # data aes(????) # var mapping geom_umap() # layer ``` --- # Posit Conf speaker Orientation, July 18, 2025: "Give a Ted Talk like Sathiya" Sathiya Manivannan, Center for Cardiovascular and Pulminary Research  --- Posit::conf(2025) Speaker Orientation ---  --- > "It's easier to type thing which are smaller, but when things are getting bigger and bigger you prefer to click on buttons and work [with a GUI]." (made a shiny wrapper for {seurat}) -- But maybe there isn't a good ggplot2 extension to handle this? ---  -- [2025-07-18 t-SNE 'labnotebook' experiment](https://evamaerey.github.io/mytidytuesday/2025-07-18-seurat_tsne_plot/seurat_tsne_plot.html) --- ## Nashville JSM Talk, "Developments in Statistical Graphics" panel, Tuesday, August 5  <!-- --- --> <!--  --> <!-- <https://phhp.ufl.edu/about/departments/biostatistics/about/faculty/> --> ---  -- [umap labnotebook experiment, 2025-08-19](https://evamaerey.github.io/mytidytuesday/2025-08-19-umap/umap.html) --- ``` r ggplot(voting_data) + # data aes(????) # var mapping geom_pca() # layer - options inside ggplot(voting_data) + # data aes(????) # var mapping geom_tsne() # layer ggplot(voting_data) + # data aes(????) # var mapping geom_umap() # layer ``` --- ### June-Sept 2024... `geom_pca_point()`  --- ``` r palmerpenguins::penguins %>% ggplot() + aes(outcome = species, pred1 = bill_length_mm, pred2 = bill_depth_mm, pred3 = flipper_length_mm) + geom_pca_point() ``` ---  --- ``` r palmerpenguins::penguins %>% ggplot() + aes(outcome = species, dims = vars(bill_length_mm, bill_depth_mm, flipper_length_mm)) + geom_pca_point() ``` --- tsne, umap - big data (lots of variables) > "It's easier to type thing which are smaller, but when things are getting bigger and bigger you prefer to click on buttons and work [with a GUI]." -- `aes(dims = vars(var1, var2, var3, var4, var5, var6)` -- `aes(dims = dims(var1:var6))` ---  --- ``` r library(ggplot2) ggplot(data = my_high_dimensional_data) + aes(dims = dims(var1:var200, var205)) + # or similar geom_reduction_technique() ``` --- ### utilities ``` r ggdims::dims_listed() ggdims::data_vars_unpack() ggdims::dims_expand() #<< expand var1:var20 ``` --- ### user-facing functions ``` r ggdims::geom_pca() ggdims::geom_tsne() ggdims::geom_umap() ``` --- # Does it work? --- # concise, consistent, 'graphical poems' ``` r library(ggdims) ggplot(unga_rcid_wide) + * aes(dims = dims(rc3:rc9147)) + * geom_pca() + aes(fill = continent) + labs(title = "PCA") -> unga_pca ``` --- # concise, consistent, 'graphical poems' ``` r library(ggdims) ggplot(unga_rcid_wide) + * aes(dims = dims(rc3:rc9147)) + * geom_tsne() + aes(fill = continent) + labs(title = "t-SNE") -> unga_tsne ``` --- # concise, consistent, 'graphical poems' ``` r library(ggdims) ggplot(unga_rcid_wide) + * aes(dims = dims(rc3:rc9147)) + * geom_umap() + aes(fill = continent) + labs(title = "UMAP") -> unga_umap ``` --- ``` r library(patchwork) unga_pca + unga_tsne + unga_umap + plot_layout(guides = "collect") + plot_annotation(title = "UN General Assembly Role Call Voting") ``` <!-- --> ---  --- How does it work? (for extenders meetup) Under the hood: ``` r ggplot(my_data) + aes(dims = dims(var1:var3, var7)) + dims_expand() + geom_umap0() ``` # 1. `dims_expand` - a. look at plot data and mapping -- - b. extract implicitly referenced variables in mapping from plot data var1:var3, var7 .... to var1, var2, var3, var7 --- # 1. `dims_expand` - c. overwrite dims mapping with expression that lists all implicitly reference vars (future implementation my be new list col var that contains all referenced vars). -- `dims = dims(var1:var3, var7)` `dims = dims_listed(var1, var2, var3, var7)` --- # 2. helper `geom_umap0` uses overwritten dim mapping for dimension reduction, taking input vars and computing x and y based on type dimred --- # 3. We can write user-facing `geom_umap()`, bundles `dims_expand()` and helper `geom_umap0()` ``` r geom_umap <- function(...){ list(dims_expand(), geom_umap0(...)) } ``` --- ### Getting: ``` r ggplot(my_data) + aes(dims = dims(var1:var10, var15)) + geom_umap() ``` --- ## Across-methods fly-over (pca, tsne, umap) -- ## Within-method fly-over? (tsne specifications, data) ---  ---  ---  --- <!-- --> --- count: false .panel1-pp2-auto[ ``` r *two_clusters ``` ] .panel2-pp2-auto[ ``` dim1 dim2 type 1 -0.5357287 -0.158254930 A 2 -0.4603359 -0.090857442 A 3 -0.3965400 -0.181554289 A 4 -0.4974581 0.106132971 A 5 -0.5729073 -0.007593519 A 6 -0.4308280 -0.045092558 A 7 -0.4197852 -0.012650899 A 8 -0.6454726 -0.008424184 A 9 -0.4513991 -0.214707887 A 10 -0.3131011 -0.020732053 A 11 -0.5635106 0.200063307 A 12 -0.6802282 0.113228751 A 13 -0.4671671 -0.027005700 A 14 -0.4593054 -0.068642035 A 15 -0.6159025 0.139260603 A 16 -0.5274706 -0.048721368 A 17 -0.4913888 0.091440641 A 18 -0.4746525 0.105385605 A 19 -0.6631743 0.134228505 A 20 -0.4540370 0.049525989 A 21 -0.5340863 -0.022069238 A 22 -0.4591989 0.068009894 A 23 -0.3724366 -0.024270429 A 24 -0.5338696 0.060912500 A 25 -0.4881712 0.066703762 A 26 -0.4479319 0.145153683 A 27 -0.4100356 0.036337216 A 28 -0.5366970 -0.029124000 A 29 -0.4113923 -0.023560498 A 30 -0.4349242 -0.025011625 A 31 -0.5059801 0.026554423 A 32 -0.3454215 -0.104671746 A 33 -0.6856915 -0.036302949 A 34 -0.4129278 -0.008105794 A 35 -0.2584186 -0.095262008 A 36 -0.5828491 0.020498686 A 37 -0.7487111 0.093079867 A 38 -0.3775321 -0.072232954 A 39 -0.5980546 -0.144940645 A 40 -0.4076318 -0.083561926 A 41 -0.3919189 -0.041298155 A 42 -0.6375004 -0.244553739 A 43 -0.6168814 -0.190197900 A 44 -0.5315187 -0.062031195 A 45 -0.5145287 -0.046462443 A 46 -0.4160438 0.197317704 A 47 -0.3931976 -0.041479515 A 48 -0.6571367 -0.046417266 A 49 -0.5226180 -0.120945937 A 50 -0.3934646 0.168484206 A 51 -0.4833463 -0.014768899 A 52 -0.5692205 0.044248820 A 53 -0.4926059 0.033500087 A 54 -0.3977887 -0.083249030 A 55 -0.2947652 -0.140350732 A 56 -0.6452856 0.094217235 A 57 -0.5158384 -0.189577055 A 58 -0.4925182 -0.087128492 A 59 -0.6553099 -0.017168386 A 60 -0.4006728 -0.222864041 A 61 -0.4146802 0.129645996 A 62 -0.4074571 -0.003922020 A 63 -0.5670353 -0.096400143 A 64 -0.4133183 -0.029236775 A 65 -0.5634832 0.127548206 A 66 -0.5671065 -0.129206785 A 67 -0.4521314 0.058483023 A 68 -0.3951574 -0.132199846 A 69 -0.2594366 0.063479376 A 70 -0.3845447 0.048875937 A 71 -0.4612730 0.029444030 A 72 -0.3541722 0.065608719 A 73 -0.3481411 -0.225318419 A 74 -0.4471973 0.108038841 A 75 -0.4235374 0.042559782 A 76 -0.4922954 0.081371883 A 77 -0.4545600 -0.053815694 A 78 -0.5857024 -0.037144865 A 79 -0.5018638 -0.038791169 A 80 -0.4938678 -0.059709975 A 81 -0.3750402 0.010759252 A 82 -0.4374003 -0.055493015 A 83 -0.5234888 0.103608133 A 84 -0.5619091 0.062872984 A 85 -0.4495126 -0.030400778 A 86 -0.3325314 0.012240477 A 87 -0.3489993 -0.092286488 A 88 -0.2913570 -0.086854253 A 89 -0.5642657 -0.017283381 A 90 -0.6731230 -0.072955045 A 91 -0.5319685 -0.099023766 A 92 -0.6504516 0.288148443 A 93 -0.6074628 0.046699775 A 94 -0.5617616 -0.007106916 A 95 -0.5260821 -0.098312730 A 96 -0.5227355 0.004991032 A 97 -0.5499129 0.163495761 A 98 -0.3925901 -0.104594754 A 99 -0.4356371 0.044225388 A 100 -0.4654224 0.070153340 A 101 -0.4187834 -0.186017354 A 102 0.5606903 0.135705951 B 103 0.4797486 -0.055312440 B 104 0.4183161 -0.081727621 B 105 0.6608417 -0.057504167 B 106 0.4696202 0.038560367 B 107 0.5108098 -0.055498693 B 108 0.4883599 -0.026284564 B 109 0.4410329 -0.001418396 B 110 0.4625084 0.079891023 B 111 0.3538451 0.028688468 B 112 0.3674274 0.035336611 B 113 0.4891274 -0.159566779 B 114 0.5566714 0.109369754 B 115 0.4966641 0.004084352 B 116 0.6231104 -0.041932167 B 117 0.5384671 -0.013715868 B 118 0.7544933 0.067691117 B 119 0.5334281 0.106882091 B 120 0.5845310 0.020439962 B 121 0.5000153 0.101844066 B 122 0.4748349 0.032864464 B 123 0.3542561 -0.067419903 B 124 0.4568676 0.172067517 B 125 0.2951672 0.181870248 B 126 0.5274469 0.034762617 B 127 0.3738903 -0.089173900 B 128 0.6319767 0.013811068 B 129 0.4758679 -0.112147293 B 130 0.2392581 0.019593448 B 131 0.5963978 0.058669812 B 132 0.5795330 0.059367882 B 133 0.4463065 0.045962269 B 134 0.4762261 -0.187948212 B 135 0.3813057 -0.082105191 B 136 0.5990435 -0.032782405 B 137 0.4256180 -0.156611906 B 138 0.6521080 -0.119362165 B 139 0.4919072 -0.058501426 B 140 0.5210185 0.140633282 B 141 0.5962070 -0.064638501 B 142 0.7162381 -0.322461265 B 143 0.5825375 0.102024799 B 144 0.4554914 -0.254290157 B 145 0.6458607 0.041606003 B 146 0.4521540 -0.015526346 B 147 0.3281885 0.076548531 B 148 0.5946617 -0.064311903 B 149 0.6148766 -0.143126072 B 150 0.5905803 0.099651771 B 151 0.4898780 0.057742666 B 152 0.3730359 0.013030242 B 153 0.3798805 0.155704161 B 154 0.4859656 -0.151298359 B 155 0.6868888 0.090086722 B 156 0.3912322 -0.043332386 B 157 0.4152529 -0.108574521 B 158 0.5099164 0.051162112 B 159 0.4358920 0.047650274 B 160 0.5475221 0.015635954 B 161 0.4009169 0.071916454 B 162 0.6612316 0.191754664 B 163 0.5222793 0.151908888 B 164 0.4604178 -0.084217285 B 165 0.6336940 0.008370056 B 166 0.5569161 -0.095242013 B 167 0.3630500 -0.046647038 B 168 0.5259380 0.011356280 B 169 0.6010525 -0.011316699 B 170 0.4744200 0.051207119 B 171 0.5730595 -0.137777461 B 172 0.6183336 0.141974631 B 173 0.5196704 -0.005082690 B 174 0.4003640 -0.068865048 B 175 0.4272597 0.005734501 B 176 0.4941395 0.148361504 B 177 0.5752577 -0.013311538 B 178 0.4736307 -0.054618941 B 179 0.3304741 -0.100005928 B 180 0.5609077 0.041335106 B 181 0.5010521 -0.023998367 B 182 0.5115131 -0.013954765 B 183 0.2969717 0.146663327 B 184 0.4235824 0.010328176 B 185 0.4767219 0.083918737 B 186 0.4321685 -0.034251659 B 187 0.5426292 0.007279260 B 188 0.5737919 -0.005723749 B 189 0.5357621 0.163366253 B 190 0.4604271 -0.030695260 B 191 0.7259624 0.046748228 B 192 0.3803555 0.044444520 B 193 0.5401534 -0.013522739 B 194 0.4126256 0.105441312 B 195 0.5047353 0.037931225 B 196 0.4851634 -0.140709581 B 197 0.4321986 -0.017397553 B 198 0.3771970 -0.037598042 B 199 0.4394506 0.049680431 B 200 0.4491798 0.061752594 B 201 0.5035890 -0.140302578 B 202 0.5967145 0.027716646 B ``` ] --- count: false .panel1-pp2-auto[ ``` r two_clusters |> * ggplot() ``` ] .panel2-pp2-auto[ <!-- --> ] --- count: false .panel1-pp2-auto[ ``` r two_clusters |> ggplot() + * aes(dims = dims(dim1:dim2)) ``` ] .panel2-pp2-auto[ <!-- --> ] --- count: false .panel1-pp2-auto[ ``` r two_clusters |> ggplot() + aes(dims = dims(dim1:dim2)) + * geom_tsne(perplexity = 2) ``` ] .panel2-pp2-auto[ <!-- --> ] --- count: false .panel1-pp2-auto[ ``` r two_clusters |> ggplot() + aes(dims = dims(dim1:dim2)) + geom_tsne(perplexity = 2) + * labs(title = "perplexity = 2") ``` ] .panel2-pp2-auto[ <!-- --> ] --- count: false .panel1-pp2-auto[ ``` r two_clusters |> ggplot() + aes(dims = dims(dim1:dim2)) + geom_tsne(perplexity = 2) + labs(title = "perplexity = 2") + * aes(fill = type) ``` ] .panel2-pp2-auto[ <!-- --> ] --- count: false .panel1-pp2-auto[ ``` r two_clusters |> ggplot() + aes(dims = dims(dim1:dim2)) + geom_tsne(perplexity = 2) + labs(title = "perplexity = 2") + aes(fill = type) -> *pp2; pp2 ``` ] .panel2-pp2-auto[ <!-- --> ] <style> .panel1-pp2-auto { color: black; width: 49%; hight: 32%; float: left; padding-left: 1%; font-size: 80% } .panel2-pp2-auto { color: black; width: 49%; hight: 32%; float: left; padding-left: 1%; font-size: 80% } .panel3-pp2-auto { color: black; width: NA%; hight: 33%; float: left; padding-left: 1%; font-size: 80% } </style> --- count: false .panel1-pp5-2[ ``` r two_clusters |> ggplot() + aes(dims = dims(dim1:dim2)) + geom_tsne(perplexity = 5) + labs(title = "perplexity = 5") + aes(fill = type) -> pp5; pp5 ``` ] .panel2-pp5-2[ <!-- --> ] --- count: false .panel1-pp5-2[ ``` r two_clusters |> ggplot() + aes(dims = dims(dim1:dim2)) + geom_tsne(perplexity = 5) + labs(title = "perplexity = 5") + aes(fill = type) -> pp5; pp5 ``` ] .panel2-pp5-2[ <!-- --> ] <style> .panel1-pp5-2 { color: black; width: 49%; hight: 32%; float: left; padding-left: 1%; font-size: 80% } .panel2-pp5-2 { color: black; width: 49%; hight: 32%; float: left; padding-left: 1%; font-size: 80% } .panel3-pp5-2 { color: black; width: NA%; hight: 33%; float: left; padding-left: 1%; font-size: 80% } </style> --- count: false .panel1-pp30-2[ ``` r two_clusters |> ggplot() + aes(dims = dims(dim1:dim2)) + geom_tsne(perplexity = 30) + labs(title = "perplexity = 30") + aes(fill = type) -> pp30; pp30 ``` ] .panel2-pp30-2[ <!-- --> ] --- count: false .panel1-pp30-2[ ``` r two_clusters |> ggplot() + aes(dims = dims(dim1:dim2)) + geom_tsne(perplexity = 30) + labs(title = "perplexity = 30") + aes(fill = type) -> pp30; pp30 ``` ] .panel2-pp30-2[ <!-- --> ] <style> .panel1-pp30-2 { color: black; width: 49%; hight: 32%; float: left; padding-left: 1%; font-size: 80% } .panel2-pp30-2 { color: black; width: 49%; hight: 32%; float: left; padding-left: 1%; font-size: 80% } .panel3-pp30-2 { color: black; width: NA%; hight: 33%; float: left; padding-left: 1%; font-size: 80% } </style> --- count: false .panel1-pp50-2[ ``` r two_clusters |> ggplot() + aes(dims = dims(dim1:dim2)) + geom_tsne(perplexity = 50) + labs(title = "perplexity = 50") + aes(fill = type) -> pp50; pp50 ``` ] .panel2-pp50-2[ <!-- --> ] --- count: false .panel1-pp50-2[ ``` r two_clusters |> ggplot() + aes(dims = dims(dim1:dim2)) + geom_tsne(perplexity = 50) + labs(title = "perplexity = 50") + aes(fill = type) -> pp50; pp50 ``` ] .panel2-pp50-2[ <!-- --> ] <style> .panel1-pp50-2 { color: black; width: 49%; hight: 32%; float: left; padding-left: 1%; font-size: 80% } .panel2-pp50-2 { color: black; width: 49%; hight: 32%; float: left; padding-left: 1%; font-size: 80% } .panel3-pp50-2 { color: black; width: NA%; hight: 33%; float: left; padding-left: 1%; font-size: 80% } </style> --- count: false .panel1-pp100-2[ ``` r two_clusters |> ggplot() + aes(dims = dims(dim1:dim2)) + geom_tsne(perplexity = 100) + labs(title = "perplexity = 100") + aes(fill = type) -> pp100; pp100 ``` ] .panel2-pp100-2[ <!-- --> ] --- count: false .panel1-pp100-2[ ``` r two_clusters |> ggplot() + aes(dims = dims(dim1:dim2)) + geom_tsne(perplexity = 100) + labs(title = "perplexity = 100") + aes(fill = type) -> pp100; pp100 ``` ] .panel2-pp100-2[ <!-- --> ] <style> .panel1-pp100-2 { color: black; width: 49%; hight: 32%; float: left; padding-left: 1%; font-size: 80% } .panel2-pp100-2 { color: black; width: 49%; hight: 32%; float: left; padding-left: 1%; font-size: 80% } .panel3-pp100-2 { color: black; width: NA%; hight: 33%; float: left; padding-left: 1%; font-size: 80% } </style> --- ``` r library(patchwork) original + pp2 + pp5 + pp30 + pp50 + pp100 ``` <!-- --> --- ``` r library(patchwork) original + pp2 + pp5 + pp30 + pp50 + pp100 & guides(fill = "none") ``` <!-- --> --- ``` r library(patchwork) original + pp2 + pp5 + pp30 + pp50 + pp100 & guides(fill = "none") & theme_ggdims() ``` <!-- --> --- ``` r library(patchwork) original + pp2 + pp5 + pp30 + pp50 + pp100 & guides(fill = "none") & theme_ggdims() -> two_clusters_5_perplexities_tsne ``` ---  --- ``` r two_clusters_5_perplexities_tsne & # contains all six of the 'hello world of tsne' specifications two_close_and_one_far #<< # change data ``` <!-- --> ---  --- ``` r two_clusters_5_perplexities_tsne & #contains all six of the 'hello world of tsne' specifications big_and_small_cluster # change data ``` <!-- --> ---  --- ``` r two_clusters_5_perplexities_tsne & # contains all six of the 'hello world of tsne' specifications random_noise & # change data aes(fill = I("navy")) ``` <!-- --> --- # Is this the right implementation? -- Maybe not -- # Are there some known fragilities -- Yes -- # Are you the right person to be writing things like `geom_umap` -- Maybe not.. --- class: inverse, middle, center Thank you! --- ### To Do #### gg side: `aes(dims = dims())` and `unpack()` implementation - ggpcp's approach (Heike)? - ggswim approach (Teun and Rich, via June C.)? -- - fundamentally, can `aes(dims = dims())` be totally compatible w/ ggplot2? -- - ggplot2 allows new vars to be computed on-the-fly in aes, but classically from vectors (be they referenced from variables data or otherwise). --- ### To do #### on the dimred side - layer_pca/layer_tsne/layer_umap requirements (e.g. Cory, Joyce for PCA) - borrow internals with tidymodels for consistency? ---  --- <style type="text/css"> .remark-code{line-height: 1.5; font-size: 70%} @media print { .has-continuation { display: block; } } code.r.hljs.remark-code{ position: relative; overflow-x: hidden; } code.r.hljs.remark-code:hover{ overflow-x:visible; width: 500px; border-style: solid; } </style>